Protein sequence data for the HA segment of all circulating A/H1N1 (seasonal and 2009 pandemic), A/H3N2, and B strains collected from human hosts between -01 were downloaded from GISAID. 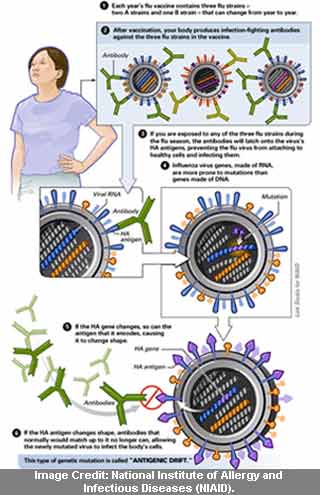
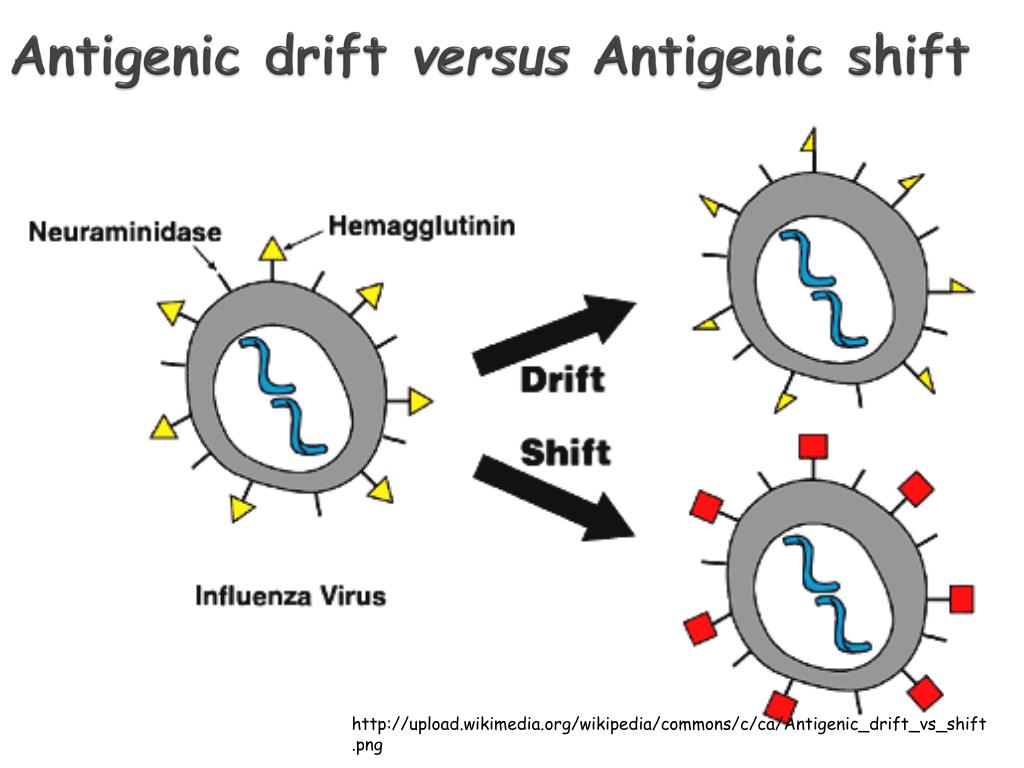
In this study, we aim to assess if a statistical relationship can be detected between the antigenic drift of seasonal influenza and the epidemiological severity of seasonal epidemics. However, to our knowledge, no studies have measured this association within a Canadian context. Previous studies have integrated sequence-based antigenic distances and epidemiological measures of severity obtained through surveillance data to link antigenic drift to seasonal severity 9, 10. Thanks to the sequencing efforts shared publicly by many participating laboratories worldwide, it is possible to infer the antigenic drift of circulating influenza viruses across many seasons using these computational methods. More recently, methods based on changes to genetic sequences have been explored as an alternative approach to antigenic cartography 6, 7, 8. Traditionally, antigenic differences between viruses have been determined experimentally using laboratory procedures such as the hemagglutination inhibition assay however, these methods are time-consuming and costly. In particular, mutations in the hemagglutinin protein, which contains antigenic sites targeted by neutralizing antibodies, can lead to immune escape and may also have implications for vaccine effectiveness 4, 5. Although epidemic influenza dynamics are complex and not well-understood, one of the major drivers of this seasonal variation is posited to be the antigenic drift of influenza viruses 3, resulting from the accumulation of mutations from one season to another. Epidemic severity varies from season to season. 
In Canada, it has been estimated that approximately 12,200 hospitalizations deaths 2 are attributed to seasonal influenza epidemics on average each year. Seasonal influenza epidemics occur globally every year, leading to a high burden of mortality and morbidity, especially in children, older adults, and individuals with underlying health conditions. Future studies may need to account for additional factors, such as co-circulation of other respiratory pathogens, population imprinting, cohort effects and environmental parameters, which may drive seasonal influenza severity. We did not find any evidence of a statistical relationship between antigenic distance and seasonal influenza severity in Canada. We performed linear regressions of our severity index with respect to the inter-seasonal antigenic distances, controlling for vaccine effectiveness. These aggregate measures of disease severity were integrated into a single seasonal severity index. We also gathered epidemiological data on cases, hospitalizations and deaths from national surveillance systems and other official sources, as well as vaccine effectiveness estimates to address potential effect modification. We obtained hemagglutinin protein sequences collected in Canada between the 020 flu seasons from GISAID and calculated Hamming distances in a sequence-based approach to estimating inter-seasonal antigenic differences. In this study, we aimed to investigate the association between the genetic drift of seasonal influenza viruses (A/H1N1, A/H3N2 and B) and the epidemiological severity of seasonal epidemics within a Canadian context. One of the major drivers of this seasonal variation is thought to be the antigenic drift of influenza viruses, resulting from the accumulation of mutations in viral surface proteins. 
Seasonal influenza epidemics circulate globally every year with varying levels of severity. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
